3 Days of Intensive Training Program for Beginners or Experienced Participants
PRECISION ONCOLOGY GENOMICS
From Tumor NGS to Therapeutic Decisions
A Next Generation Sequencing Data Analysis Hands-On Program to Analyze Genomic Data for Precision Oncology
Highlights of the Course
- Intensive Learning
- Hands-On Practice
- Case Studies
- Problem Solving
- Systematic Framework
- Advance Analysis (Basic to Advance)
- Detailed Outline
- Offline Training and Q&A Sessions
- Session Recording for Practice Later
ABOUT THE MASTERCLASS
The Precision Oncology Genomics Workshop addresses a critical gap in clinical genomics education. Many professionals have consumed extensive online resources—YouTube tutorials, blog posts, research articles—yet remain uncertain how to implement tumor NGS analysis or translate genomic findings into clinical decisions for their institutions.
Conventional courses often deliver only theoretical foundations and industry terminology without practical, hands-on application. This workshop dispels common misconceptions about clinical genomics interpretation, variant classification, and multi-omics integration that hinder effective precision medicine adoption.
Participants move beyond foundational concepts to advanced applications: somatic pipeline implementation, AI-augmented variant prioritization, and integration of RNA-seq and proteomics data—equipping you with deployment-ready workflows that directly advance your research publications, institutional genomics programs, and career trajectory in precision oncology.
That's Why We Created 3 Day Precision Oncology Genomics Masterclass
The Precision Oncology Genomics Masterclass was conceptualized to provide a comprehensive walkthrough of the complete clinical genomics workflow—encompassing NGS data analysis, variant annotation, multi-omics integration, AI-augmented interpretation, and therapeutic decision-making. By integrating these interconnected technologies into a unified pipeline, participants gain a holistic perspective of how precision oncology functions in real-world clinical settings. Throughout the 3-day curriculum, real-world cancer case studies and institutional implementation scenarios anchor theoretical concepts in practical application, enabling participants to deploy complete, validated workflows within their hospitals and research centers.
This Is A 100% Practical Training Program Where You Will Learn By DOING THE STEPS And Have Your Own Dashboard To Showcase Your Results.
DIFFERENT TECHNOLOGIES INCLUDED IN THE MASTERCLASS
A number of different steps are included in the pipeline which are all together part of different technology only. So, below we have tried to list down the different technologies used to form the pipeline for carrying out the analysis.

High-throughput DNA sequencing technology enabling comprehensive analysis of tumor genomes, detecting somatic mutations, structural variants, and chromosomal abnormalities with clinical precision and cost-effectiveness.

Computational workflows processing raw sequencing data through quality control, alignment, and variant detection algorithms to identify pathogenic mutations, distinguish somatic from germline variants, and generate standardized variant calls.

Systematic annotation of detected variants using genomic databases (ClinVar, COSMIC, OncoKB), followed by evidence-based classification using ACMG/AMP/ESMO guidelines to determine clinical actionability and therapeutic relevance.
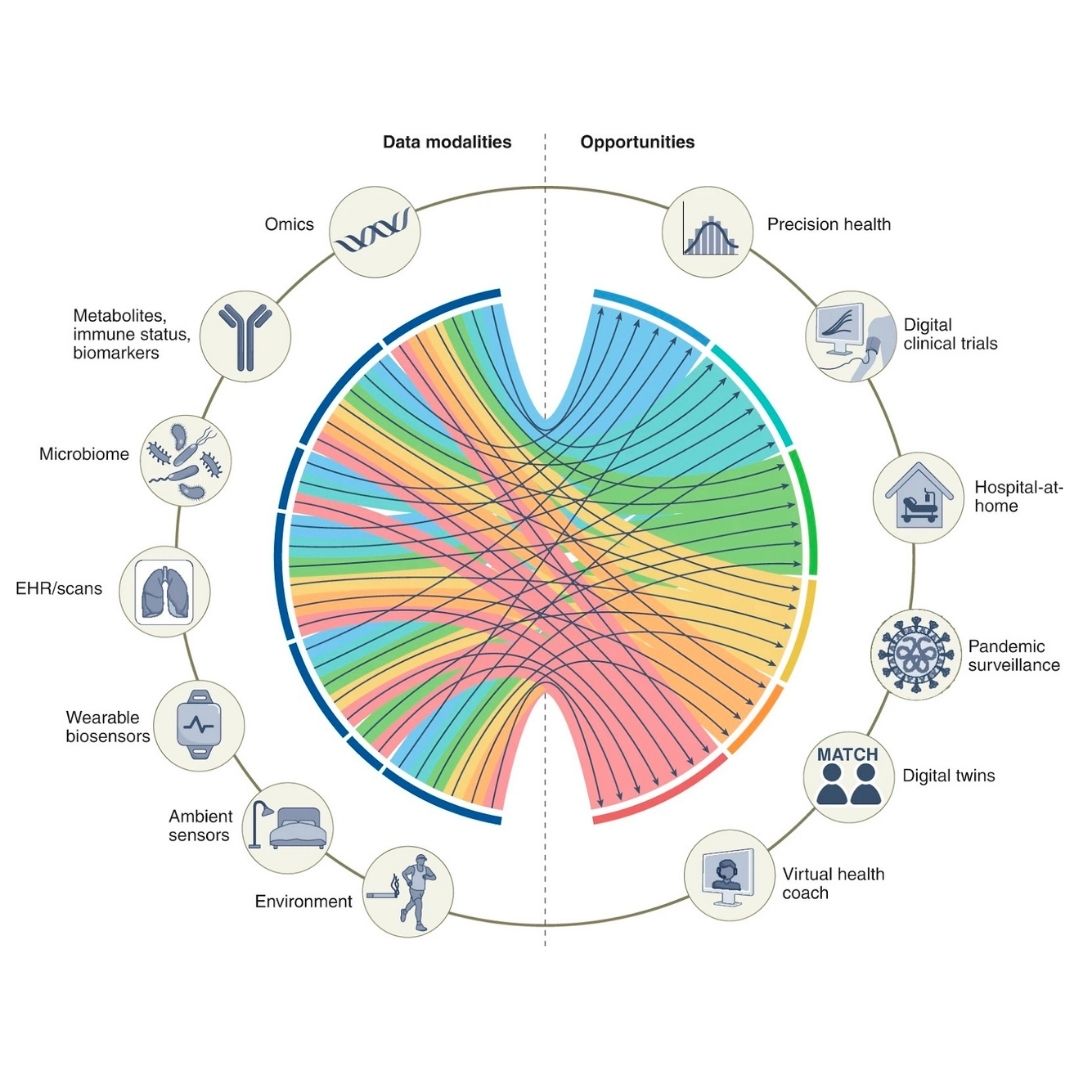
Integration of complementary genomic data—RNA-seq expression profiles, proteomics, methylation patterns—to provide comprehensive molecular profiling for improved biomarker discovery, patient stratification, and treatment response prediction.

Advanced algorithms for feature engineering, variant prioritization, and risk stratification from genomic and clinical data, enabling intelligent filtering of actionable mutations and prediction of treatment outcomes with enhanced clinical relevance.

Structured generation of hospital-ready genomics reports translating complex molecular findings into clinician-friendly summaries, therapy recommendations, and tumor board documentation aligned with regulatory standards and institutional requirements.
Mentor for the Masterclass

Co-Founder & Director Research
He is the Co-Founder & Director (Operations) EdGene BioMed Pvt Ltd, Gurugram.
His specializations include Chemoinformatics, Drug Design, Molecular Modelling Dynamics & Simulation, Next Generation Sequencing Data Analysis, T Cell Repertoire Sequencing.
He was awarded as Transformational Entrepreneur award in 2016 by Versys Labs
(Virginia Tech University) in Bangalore.
He has an experience of 8 Years in the field of Molecular Docking & MD Simulation and has conducted several National & International training programs.
Detailed Masterclass Course Module
Different modules have been layed out which we will be covering over the period of 12 weeks to learn the process better and get hands-on learning experience working on them.
Day 0 - Pre-Masterclass Preparatory Course (Pre-Recorded)

1. Introduction to important terminologies used in the process
2. Introduction to the tools used in the process
3. Running the process in GUI based tool for better understanding of the process
4. Preparing your systems for the masterclass
5. Downloading the data for the masterclass
Day 1 (March 22, 2026) - Foundations & Data Infrastructure

1. Precision oncology landscape in Middle East healthcare,
2. Tumor biology: somatic vs. germline mutations
3. NGS technologies: panel, exome, and whole-genome sequencing
4. Sample types and pre-analytical considerations (FFPE, fresh tissue)
5. Quality metrics and acceptance criteria
6. Introduction to bioinformatics pipelines

1. FASTQ quality control and preprocessing
2. Reference genome selection and alignment strategies
3. Somatic variant calling algorithms and best practices
4. Distinguishing true variants from sequencing artifacts
5. Depth of coverage analysis and sensitivity thresholds
6. Hands-on: Process real tumor datasets end-to-end
7. Practical exercise: Quality assessment checkpoints
Day 2 (March 23, 2026) - Clinical Interpretation & Actionable Insights

1. Annotation databases: ClinVar, COSMIC, CIViC, OncoKB
2. ACMG/AMP/ESMO guidelines for variant interpretation
3. Functional prediction tools and conservation analysis
4. Tier classification: pathogenic, likely pathogenic, VUS, benign
5. Population frequency databases and allele filtering
6. Hands-on: Annotate and classify variants from case studies

1. Somatic mutation burden and tumor mutational load (TMB)
2. Copy number variation (CNV) analysis and clinical significance
3. Microsatellite instability (MSI) detection and interpretation
4. Fusion gene identification and therapeutic implications
5. Homologous recombination deficiency (HRD) assessment
6. Case-based interpretation: Breast, lung, colorectal, hematologic malignancies
Day 3 (March 24, 2026) - Advanced Integration & Clinical Implementation

1. RNA-seq integration for expression-based biomarkers
2. Proteomics data interpretation and pathway analysis
3. AI/machine learning for variant triage and prioritization
4. Risk stratification models and survival prediction
5. Feature engineering from genomic and clinical data
6. Reproducibility, validation, and publication guidelines
7. Hands-on: Build and validate an ML-based variant filter

1. Clinical genomics report structure and content requirements
2. Physician-friendly interpretation and recommendations
3. Therapy matching: approved drugs, trials, compassionate use
4. Tumor board presentation and multidisciplinary discussion
5. Institutional workflow integration possibility
6. Concluding the Masterclass
Daily Learning Objectives of Topics Covered in the Masterclass
Below we have tried to outline the complete process we are going to cover in this bootcamp so that you can get a clear idea of what we are going to do and in what sequence, to help you better make the decision.
Participants understand tumor genomics concepts and can execute complete NGS data processing pipelines from raw sequencing data to variant calls.
Participants master variant classification, evidence grading, and can generate clinically justified interpretation reports aligned with international guidelines.
Participants integrate advanced analytics, deploy AI tools, and create hospital-ready genomics workflows ready for immediate institutional implementation.
MASTERCLASS TAKEAWAY
Apart from the topics mentioned above there are a few extra things which you can take away from this bootcamp, which will be adding more value to your work.
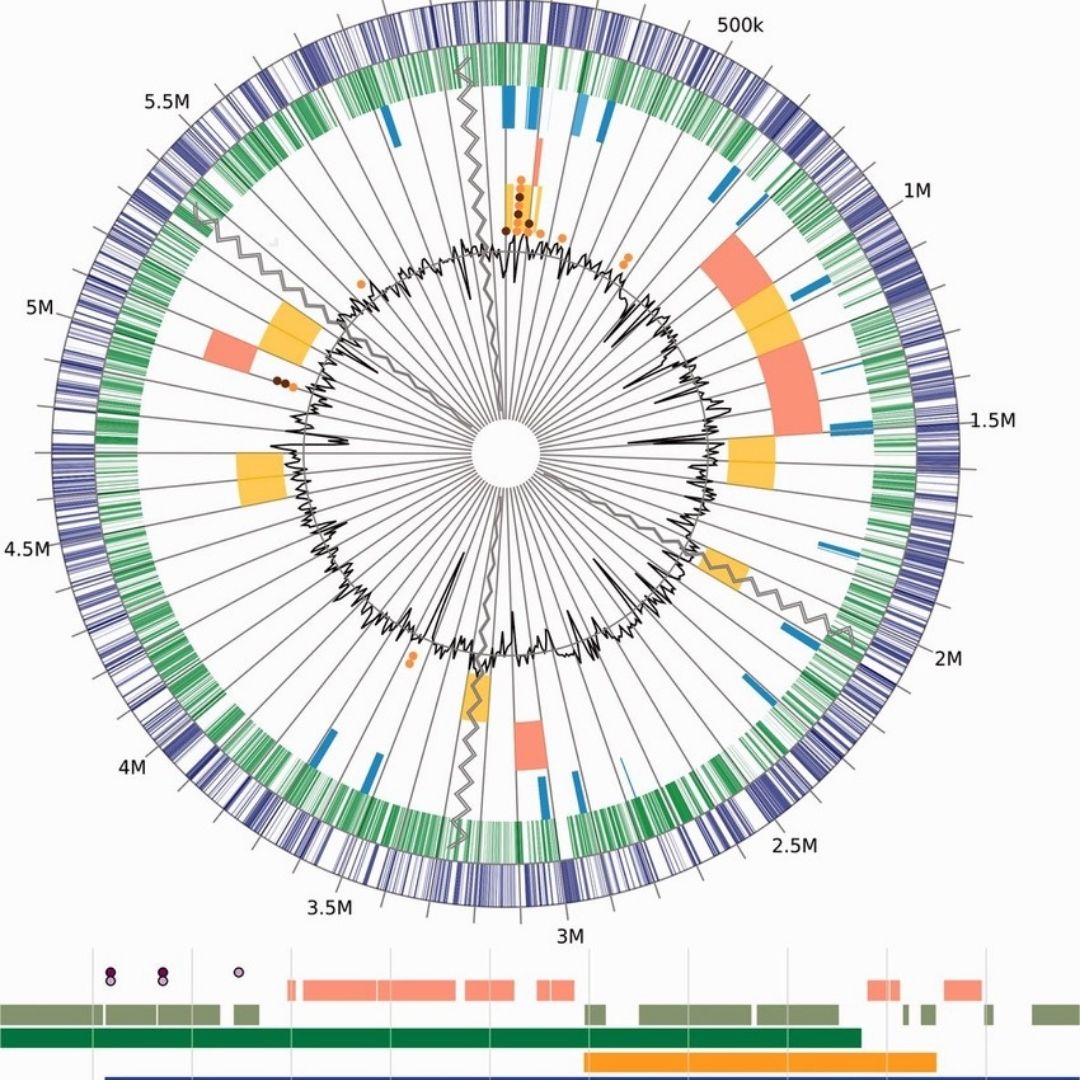
Participants gain the ability to independently run and troubleshoot a complete somatic NGS pipeline—from FASTQ QC and alignment to high-confidence variant calling—using real tumor datasets, ensuring they can reproducibly generate clinically robust genomic outputs suitable for downstream interpretation and institutional deployment.
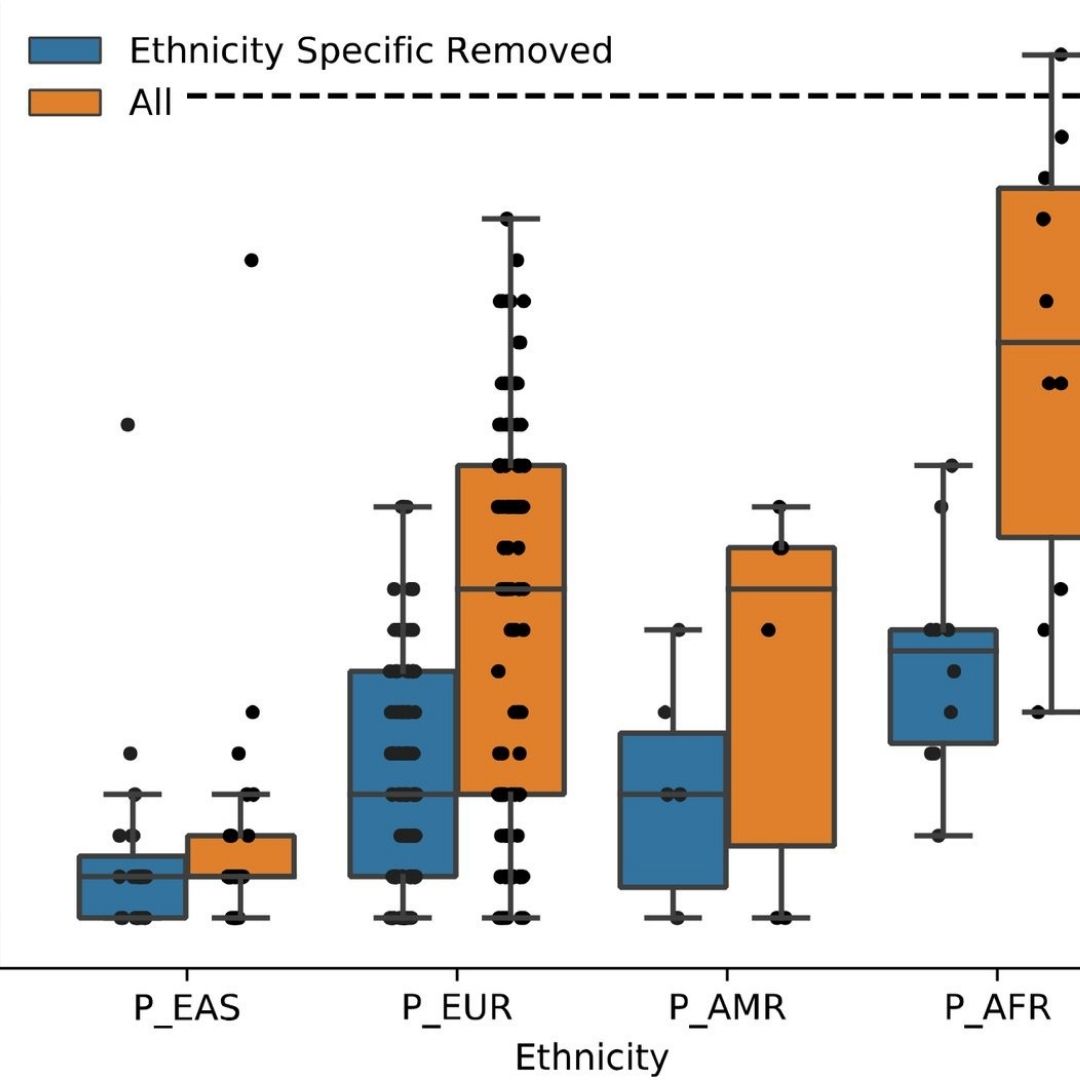
Participants develop practical skills in calculating and interpreting TMB, MSI status, CNVs, and key oncogenic fusions, understanding how these features inform immunotherapy eligibility, targeted therapy selection, and risk stratification across multiple tumor types, including lung, breast, colorectal, and hematologic cancers.
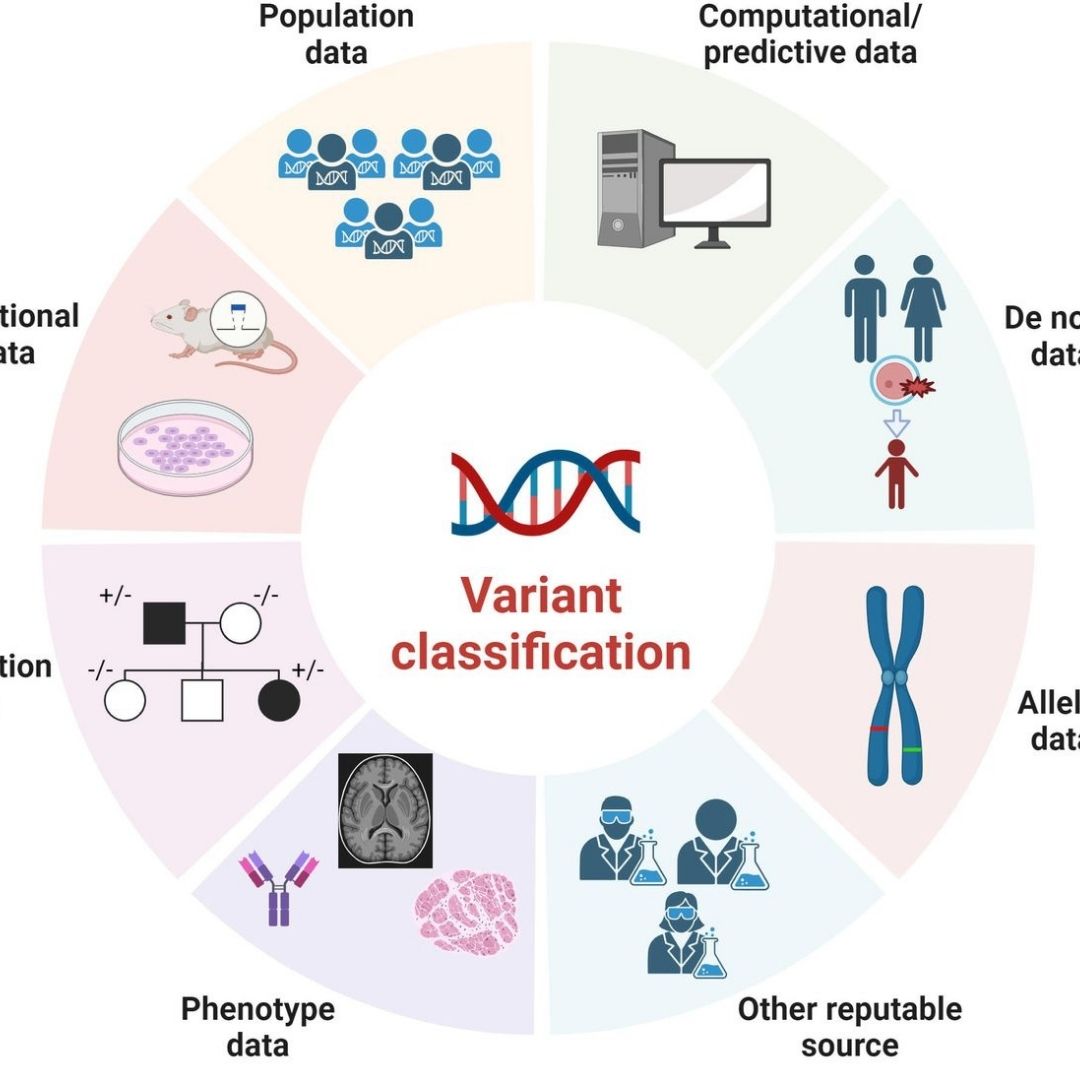
Attendees learn to annotate and classify variants using major oncology databases and ACMG, AMP, ESMO frameworks, translating complex genomic findings into tiered, evidence-backed conclusions and therapy options that can be confidently presented to tumor boards and incorporated into multidisciplinary clinical decision.
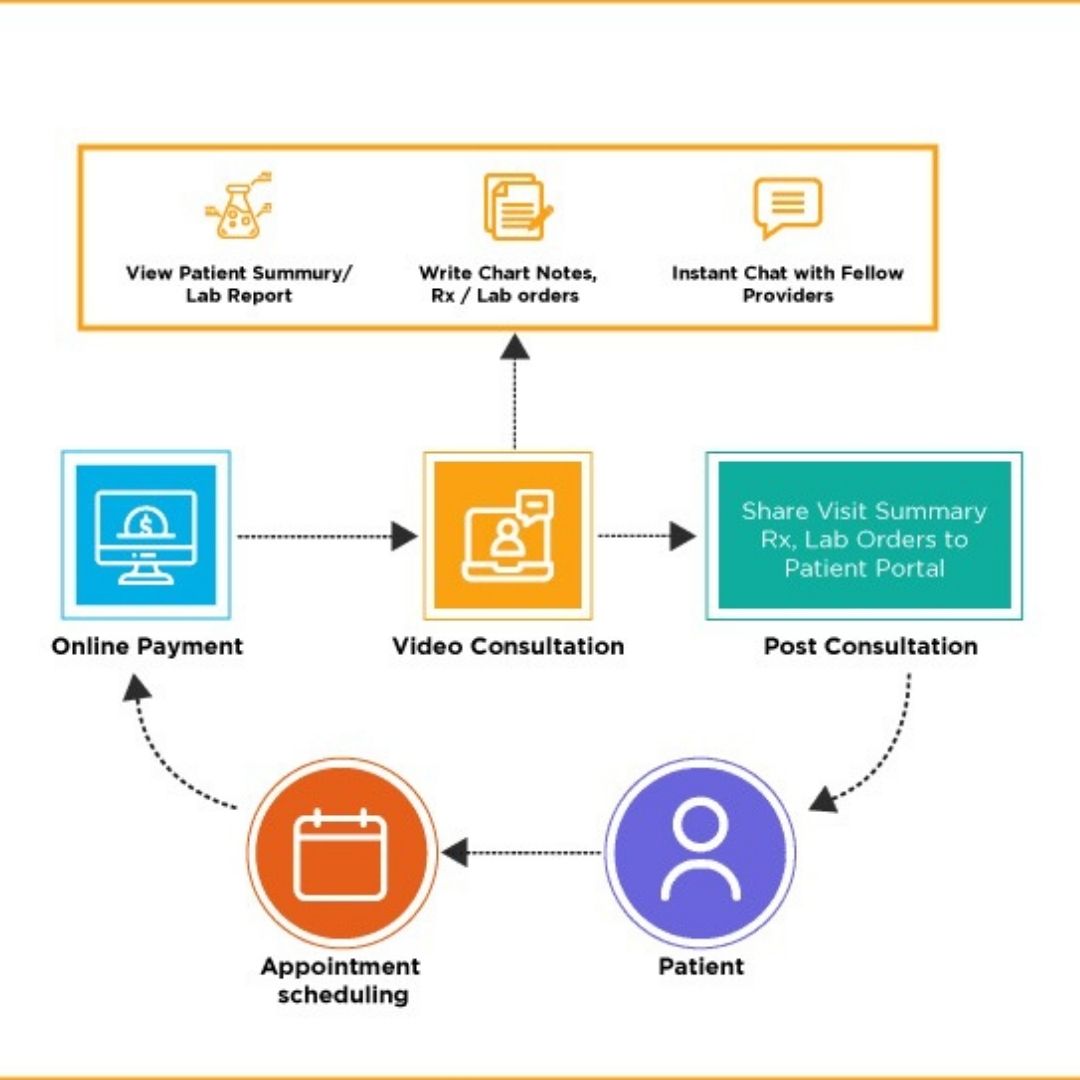
By the end, each participant assembles a complete, documented workflow—including SOPs, report templates, and integration guidelines—that can be adapted for their own hospital or lab, enabling rapid, compliant deployment of precision oncology genomics services within existing clinical and informatics infrastructures.
MASTERCLASS OUTCOMES
Let’s look into the expected outcomes of attending the Bootcamp and how it will benefit the participants to learn & Execute the process on the their own without any external help.
Able to set up, run, and document a complete tumor NGS pipeline (QC, alignment, variant calling, basic filtering) that can be validated and transferred into institutional or lab workflows.
Able to compute and interpret TMB, MSI, CNVs, and key fusions, and explain their implications for immunotherapy and targeted treatment decisions across common solid and hematologic tumors.
Can design structured, clinician-friendly reports that summarize key findings, actionability, limitations, and recommended next steps, aligned with international reporting practices for tumor boards and oncology teams.
Capable of annotating variants with major oncology databases and applying ACMG/AMP-style frameworks to classify somatic variants and justify clinical relevance in clear, evidence-supported language.
Understands how to integrate RNA-seq and other omics with DNA data and apply basic ML-driven triage concepts to prioritize variants and refine risk or response predictions.
Leaves with a clear, stepwise plan—covering technology, people, processes, and documentation—to initiate or strengthen a precision oncology genomics workflow in their own hospital, lab, or research unit.
PARTICIPANTS FEEDBACK
Below we have some of the previous participant’s feedback what they think about our training programs.
Hey all, Im on the last day of the Molecular Dynamics Simulation workshop by EdGene BioMed and believe me its a great workshop for anyone from novice to an intermediate to understand the process and conecpts in detail. I would like to extend my heartfelt thanks to the tutor, Yash Sir and the organisation!
Thank you so much
I am really glad to join this 10days workshop.
The hands-on training of MD simulations using installation of gromacs is really a wonderful experience.
The dedication and hardwork of Yash and his team is Highly appreciated who made us understand even minor details of the process.
Looking forward to do more work with the team.
Training on MD Simulation with GROMACS & MMPBSA Analysis was an excellent experience for me as I was the beginner in this field but due excellent explanation by Yash Srivastava sir I felt it easy to learn whole process. Workshop was very interactive, I appreciate that i learned a lot in a very short time they included almost all type of information about docking.
BOOK YOUR SEAT
To understand and discuss the details of the course you can book a free 30 Minutes quick meeting with us which will help you make better informed decision to take this course and get the most out of it.
Contact Us
We understand that you may have some questions before you make the payment for the course. Feel free to get in contact with us through the below given options.
Feel free to drop us a text on
For More Contact Information
